This C++ source code implements Smith Waterman Algorithm with affine gap penalties. It requires at least one blank line between the two sequences. Ignores input lines with non-alphabetical characters.
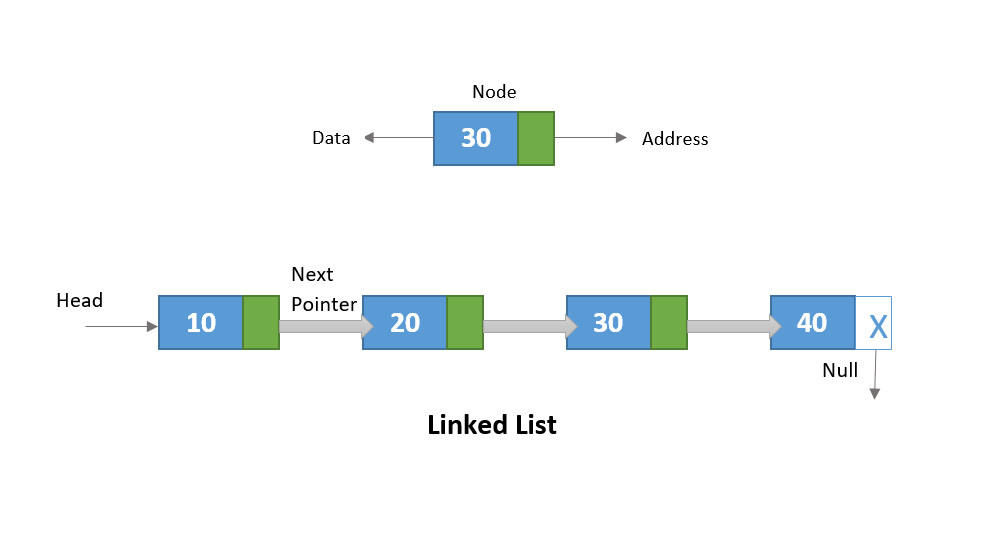
The Smith-Waterman algorithm is used for comparing two sequences, typically biological sequences like DNA, RNA, or proteins. It finds the local similarities between the sequences, identifying regions where they match or align.
Imagine you have two sequences of characters (e.g., A, C, G, T for DNA). The algorithm looks for regions in these sequences where they align well, taking into account matches, mismatches, and gaps.
The algorithm produces an optimal local alignment of the two sequences which shows the regions where they match and any gaps that are introduced to achieve this alignment. The final alignment score reflects the similarity between the aligned regions.
#include <iostream>
#include <string>
#include <cctype>
#include <algorithm>
#include <locale>
using namespace std;
const double LARGE_NUMBER = 65536.;
const double GAP_OPENING_COST = 10.;
const double GAP_EXTENSION_COST = .1;
const double NEW_GAP_COST = GAP_OPENING_COST + GAP_EXTENSION_COST;
const signed char BLOSUM[][25] = { // the blosum 62 scoring matrix
{4, 0, 0, -2, -1, -2, 0, -2, -1, 0, -1, -1, // A
-1, -2, 0, -1, -1, -1, 1, 0, 0, 0, -3, 0, -2},
{0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0},
{0, 0, 9, -3, -4, -2, -3, -3, -1, 0, -3, -1, // C
-1, -3, 0, -3, -3, -3, -1, -1, 0, -1, -2, 0, -2},
{-2, 0, -3, 6, 2, -3, -1, -1, -3, 0, -1, -4, // D
-3, 1, 0, -1, 0, -2, 0, -1, 0, -3, -4, 0, -3},
{-1, 0, -4, 2, 5, -3, -2, 0, -3, 0, 1, -3, // E
-2, 0, 0, -1, 2, 0, 0, -1, 0, -2, -3, 0, -2},
{-2, 0, -2, -3, -3, 6, -3, -1, 0, 0, -3, 0, // F
0, -3, 0, -4, -3, -3, -2, -2, 0, -1, 1, 0, 3},
{0, 0, -3, -1, -2, -3, 6, -2, -4, 0, -2, -4, // G
-3, 0, 0, -2, -2, -2, 0, -2, 0, -3, -2, 0, -3},
{-2, 0, -3, -1, 0, -1, -2, 8, -3, 0, -1, -3, // H
-2, 1, 0, -2, 0, 0, -1, -2, 0, -3, -2, 0, 2},
{-1, 0, -1, -3, -3, 0, -4, -3, 4, 0, -3, 2, // I
1, -3, 0, -3, -3, -3, -2, -1, 0, 3, -3, 0, -1},
{0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0},
{-1, 0, -3, -1, 1, -3, -2, -1, -3, 0, 5, -2, // K
-1, 0, 0, -1, 1, 2, 0, -1, 0, -2, -3, 0, -2},
{-1, 0, -1, -4, -3, 0, -4, -3, 2, 0, -2, 4, // L
2, -3, 0, -3, -2, -2, -2, -1, 0, 1, -2, 0, -1},
{-1, 0, -1, -3, -2, 0, -3, -2, 1, 0, -1, 2, // M
5, -2, 0, -2, 0, -1, -1, -1, 0, 1, -1, 0, -1},
{-2, 0, -3, 1, 0, -3, 0, 1, -3, 0, 0, -3, // N
-2, 6, 0, -2, 0, 0, 1, 0, 0, -3, -4, 0, -2},
{0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0},
{-1, 0, -3, -1, -1, -4, -2, -2, -3, 0, -1, -3, // P
-2, -2, 0, 7, -1, -2, -1, -1, 0, -2, -4, 0, -3},
{-1, 0, -3, 0, 2, -3, -2, 0, -3, 0, 1, -2, // Q
0, 0, 0, -1, 5, 1, 0, -1, 0, -2, -2, 0, -1},
{-1, 0, -3, -2, 0, -3, -2, 0, -3, 0, 2, -2, // R
-1, 0, 0, -2, 1, 5, -1, -1, 0, -3, -3, 0, -2},
{1, 0, -1, 0, 0, -2, 0, -1, -2, 0, 0, -2, // S
-1, 1, 0, -1, 0, -1, 4, 1, 0, -2, -3, 0, -2},
{0, 0, -1, -1, -1, -2, -2, -2, -1, 0, -1, -1, // T
-1, 0, 0, -1, -1, -1, 1, 5, 0, 0, -2, 0, -2},
{0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0},
{0, 0, -1, -3, -2, -1, -3, -3, 3, 0, -2, 1, // V
1, -3, 0, -2, -2, -3, -2, 0, 0, 4, -3, 0, -1},
{-3, 0, -2, -4, -3, 1, -2, -2, -3, 0, -3, -2, // W
-1, -4, 0, -4, -2, -3, -3, -2, 0, -3, 11, 0, 2},
{0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0},
{-2, 0, -2, -3, -2, 3, -3, 2, -1, 0, -2, -1, // Y
-1, -2, 0, -3, -1, -2, -2, -2, 0, -1, 2, 0, 7}
};
// trim from start (in place)
inline void ltrim(std::string& s) {
s.erase(s.begin(), std::find_if(s.begin(), s.end(), [](unsigned char ch) {
return !std::isspace(ch);
}));
}
// trim from end (in place)
inline void rtrim(std::string& s) {
s.erase(std::find_if(s.rbegin(), s.rend(), [](unsigned char ch) {
return !std::isspace(ch);
}).base(), s.end());
}
// trim from both ends (in place)
inline void trim(std::string& s) {
rtrim(s);
ltrim(s);
}
template<typename T>
class Array2D {
public:
int rows;
int cols;
T** data;
Array2D(int rows, int cols) : rows(rows), cols(cols) {
data = new T * [rows];
for (int i = 0; i < rows; ++i) {
data[i] = new T[cols]();
}
}
~Array2D() {
for (int i = 0; i < rows; ++i) {
delete[] data[i];
}
delete[] data;
}
T** getData() const {
return data;
}
// Overload the subscript operator for convenient access
T* operator[](int index) {
return data[index];
}
// Function to return the number of rows (height)
int height() const {
return rows;
}
// Function to return the number of columns (width)
int width() const {
return cols;
}
};
void read(string& sequence)
{
string line;
while (getline(cin, line)) {
trim(line);
if (line.empty()) {
return;
}
for (int i = 0, n = line.length(); i < n; i++) {
if (!isalpha(line[i] = toupper(line[i]))) {
if (!sequence.empty()) {
return;
}
line = "";
break;
}
}
sequence += line;
}
}
double max(double x, double y)
{
return x > y ? x : y;
}
double max(double x, double y, double z)
{
return x > y ? max(x, z) : max(y, z);
}
double alignment(string& s1, string& s2)
{
int n = s1.length() + 1, m = s2.length() + 1, i, j;
Array2D<double> r(n, m), t(n, m), s(n, m);
//====
// initialization
r[0][0] = t[0][0] = s[0][0] = 0;
for (i = 1; i < n; i++) {
r[i][0] = -LARGE_NUMBER;
s[i][0] = t[i][0] = -GAP_OPENING_COST - i * GAP_EXTENSION_COST;
}
for (j = 1; j < m; j++) {
t[0][j] = -LARGE_NUMBER;
s[0][j] = r[0][j] = -GAP_OPENING_COST - j * GAP_EXTENSION_COST;
}
//====
// Smith-Waterman with affine gap costs
for (i = 1; i < n; i++) {
for (j = 1; j < m; j++) {
r[i][j] =
max(r[i][j - 1] - GAP_EXTENSION_COST, s[i][j - 1] - NEW_GAP_COST);
t[i][j] =
max(t[i - 1][j] - GAP_EXTENSION_COST, s[i - 1][j] - NEW_GAP_COST);
s[i][j] =
max(
s[i - 1][j - 1] + BLOSUM[s1[i - 1] - 'A'][s2[j - 1] - 'A'],
r[i][j], t[i][j]
);
}
}
//====
// back tracking
i = n - 1, j = m - 1;
while (i > 0 || j > 0) {
if (s[i][j] == r[i][j]) {
s1.insert(i, 1, '-');
j--;
}
else if (s[i][j] == t[i][j]) {
s2.insert(j, 1, '-');
i--;
}
else {
i--, j--;
}
}
//====
// final score
return s[s.height() - 1][s.width() - 1];
}
int main()
{
string sequence1, sequence2;
read(sequence1), read(sequence2);
double score = alignment(sequence1, sequence2);
cout << sequence1 << "\n\n" << sequence2 << "\n\nScore: " << score << endl;
return 0;
}